How does our Virtuoso Sponger, URIBurner Service, and OpenLink Structure Data Sniffer interoperate en route to simplifying progressive LOD Cloud KnowledgeGraph integration from any HTML doc?
Here is a demonstration in three simple steps :
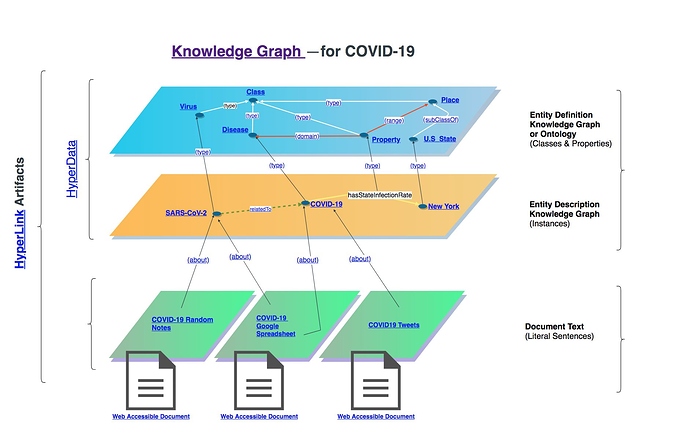
Goal: Human and machine readable variant of this: Knowledge Graph - COVID-19
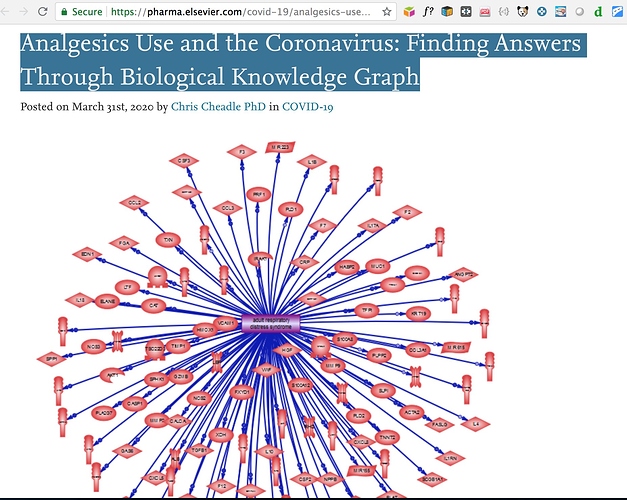
Step 1: The Original Document
Analgesics Use and the Coronavirus: Finding Answers Through Biological Knowledge Graph
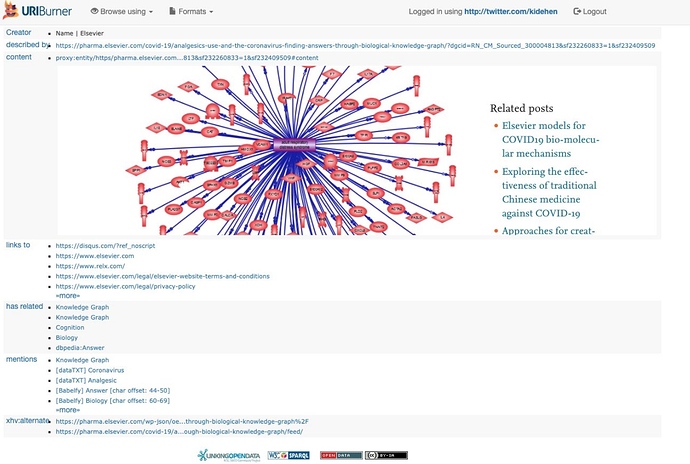
Step 2: Using the Document URL https://pharma.elsevier.com/covid-19/analgesics-use-and-the-coronavirus-finding-answers-through-biological-knowledge-graph/?dgcid=RN_CM_Sourced_300004813&sf232260833=1&sf232409509 Sponge (i.e., Extract, Transform, and Load to the LOD Cloud) via our URIBurner Service.
Sponging, as described above, takes the form of a URIBurner-specific URL: About: https://pharma.elsevier.com/covid-19/a...d_300004813 which triggers an processing pipeline that returns a description of the document, named entities, and entity relationship types that’s further enhanced via named entity recognition and extraction services looked-up from the LOD Cloud Knowledge Graph.
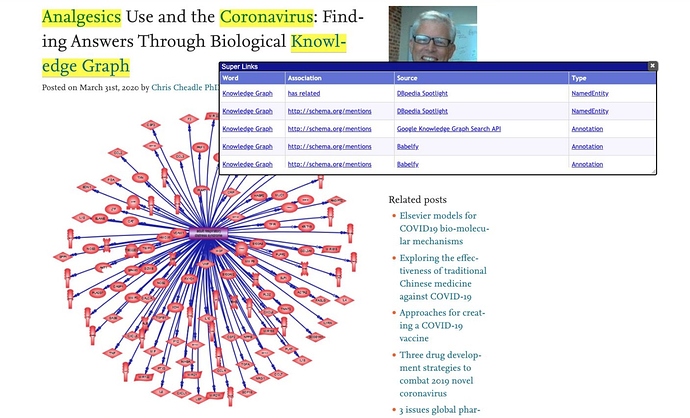
Step 3: Finally, using the OpenLink Structure Data Sniffer you can bring it all together via its Super Links feature that highlights keywords and phrases associated with entity matches; and then applies links to these highlights so that when clicked you are presented with a table of entity relationship types identified by hyperlinks that resolve to their respective descriptions. Just like a dictionary on-request.
These entry points, which are identifiers that exploit Linked Data principles, provide LOD Cloud Knowledge Graph exploration launch-points for the duration of your OSDS session.
The screenshot below depicts a COVID-19 Knowledge Graph launch-point generated from the original document that’s readable and explorable by a human or machine (e.g., software).
Conclusion
This post has demonstrated a “deceptively simple” example of the power of Linked Data and the progressively updated LOD Cloud Knowledge Graph, with an emphasis of COVID-19 research.
Related Links