There is no such thing as a “silver bullet” when dealing with the complexity associated with data access, integration, and management. In this post, I will be covering deeper change-sensitive bridging between COVID-19 and SARS-CoV-2 data sources and the Linked Open Data (LOD) Cloud Knowledge Graph.
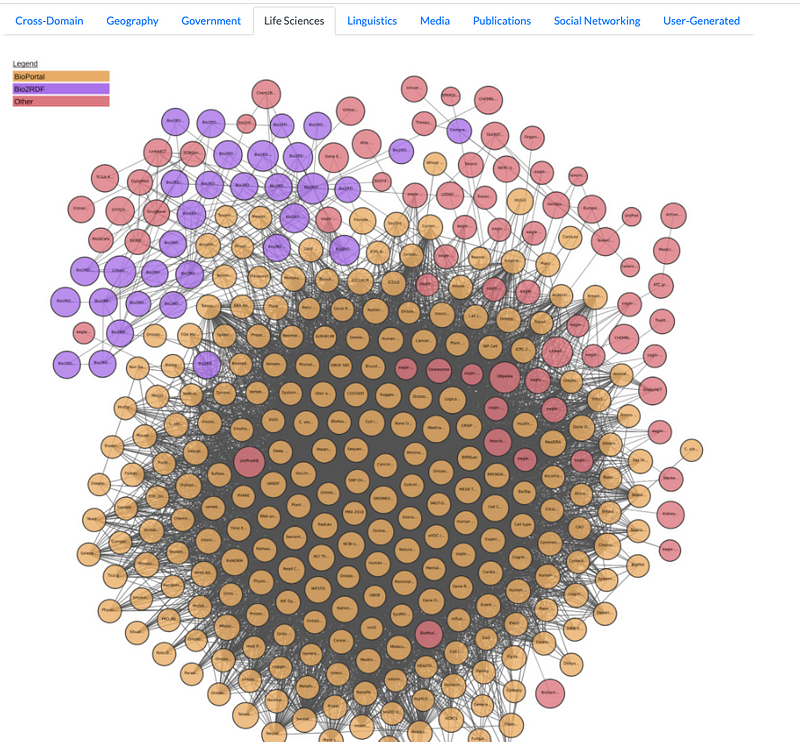
Healthcare and Life Sciences Segment of the LOD Cloud Knowledge Graph
Situation Analysis
Initially, I looked to the functionality of shared Google Spreadsheets for identifying and enhancing important COVID-19 data sources published in Open Data form. This approach enabled exploitation of the fundamental power that spreadsheets bring to bear with regards to data structure analysis and manipulation e.g., providing a highly productive way evaluate LOD Cloud Knowledge integration approaches using DBpedia identifiers for Countries, Provinces (States), and Counties.
Ultimately, Nextstrain and Johns Hopkins — two important data sources used in my Data Wrangling exercises — introduced and exposed spreadsheet limitations that I address in this post.
Problem
Nextstrain SARS-Cov-2 specimen submission tracking dataset
The is a single CSV document that has grown in size that exceeds the storage capacity of a single Google Spreadsheet i.e., all attempts to replenish content using the ImportData(“{CSV-Document-Location}”) function fail (you can click HEREto see the problem in our Master Google Spreadsheet for COVID-19 Data Sources)
Johns Hopkins Daily Reports
This collection of daily CSV documents would never have been viable for spreadsheet tracking based on the sheer number of spreadsheet tabs that would need to have been both created and then tracked.
Solution
Use the data-wrangling templates, established in the spreadsheets associated with each problematic data source, to create a DBMS processing pipeline that achieves the same goal of progressively updating the LOD Cloud Knowledge Graph without imposing anything — beyond the requirement for data to be published in Open Data form.
How
Using Virtuoso’s Technology Stack, which includes File System, Content Crawling, Content Transformation, and Content Loading functionality, here’s how we solve this challenging problem.
Nextstrain Solution
- Write a SPARQL query that targets the Github URL of the Nextstrain Metadata CSV document; include the cache-invalidation pragma ‘define get:soft “soft”’ in the query preamble — which invokes DBMS updates only if the source document has changed since the time of query invocation.
- Share the SPARQL Query via social-media outlets so that anyone can trigger a refresh, as long as they are willing to authenticate since this is a privileged activity configured in the Attribute-based Access Controls (ABAC ACLs) that underlie the URIBurner instance.
Johns Hopkins Daily Reports Solution
- Create a local fork of the Github Project.
- Create a shell script for copying the CSV document collection to a local folder situated under DB directory of your Virtuoso instance (e.g., /opt/virtuoso/database/covid-19-csv-files/), using curl or wget
- Add the shell script to the OS provided Event Scheduler e.g., crontab on Linux and Unix systems
- Create a Custom WebDAV Folder of Type “Host File FS” (meaning Host File System) that references the folder created in the prior step
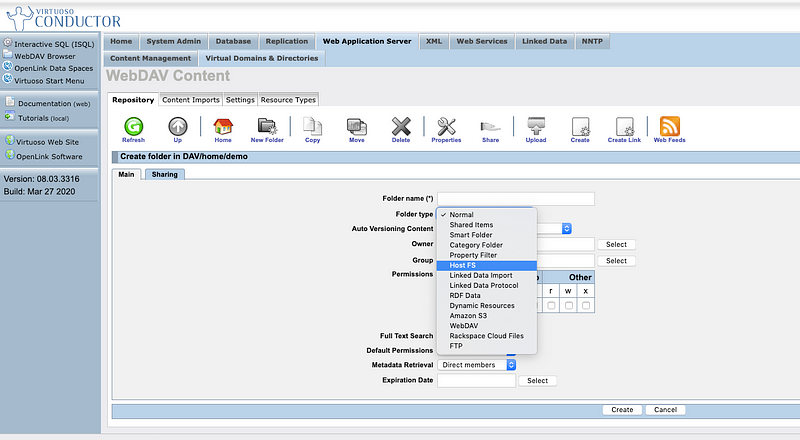
Custom WebDAV Folder Types (a/k/a Dynamic Extension Types [DETs])
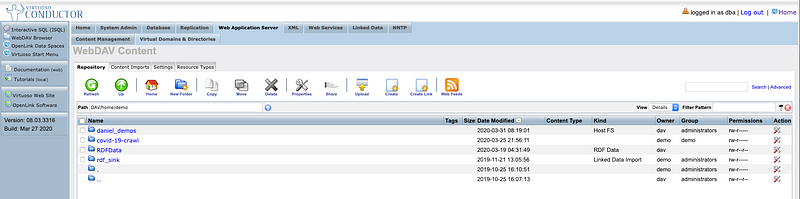
File System Folder integrated into Virtuoso’s DBMS-hosted File System that’s HTTP & WebDAV-accessible
- From the instance of Virtuoso where you seek to extract, load, and transform the CSV doc content into RDF, create a Web Content Crawler Job targeting the HTTP/WebDAV accessible folder (created in the prior step) that also enables Sponger Cartridge for CSV transformation invocation as part of the processing pipline — this includes designating a destination Named Graph for the generated RDF statements.
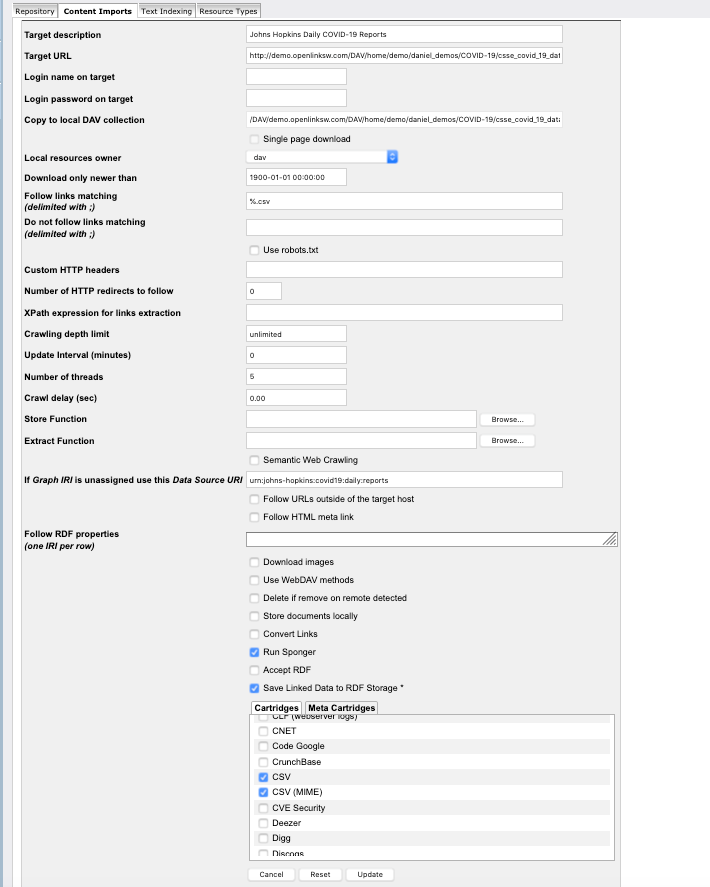
Virtuoso Crawler Job (On URIBurner Instance) Targeting a WebDAV collection exposed by another Virtuoso Instance (Demo Server)
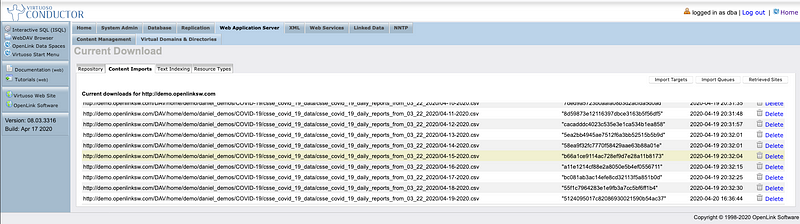
Virtuoso Crawler Dashboard Listing Documents Crawled
- Optionally, you can also create additional RDF enhancements using SPARQL INSERT statements via a Stored Procedure added to Virtuoso’s Even Scheduler
- Write a SPARQL Query targeting the Named Graph designated in the prior step and share it via social-media outlets.
Knowledge Graph Enhancements
Courtesy of SPARQL INSERT statements, you can further enhance the knowledge and presentation aesthetics of the generated RDF data with regards to:
- Entity Labels
- Entity Type (Class) Labels
- Entity Relationship Type (Properties) Labels
- Entity Relationship Type Semantics — e.g., indicating equivalence, inversion, subsumption
- Meshing Entity Identifiers with other external data sources (e.g., Bing’s change-sensitive COVID19 dashboard, Nextstrain’s highly visual specimen provenance information, and powerful taxonomy pages from the U.S. National Institutes of Health (NIH).
The scripts for achieving these tasks are part of our new COVID-19 specific github repository.
- Nextstrain Scrips
- Johns Hopkins Scripts
- General Script comprising various Reasoning and Inference Rules covering equivalence and labeling aesthetics.
SPARQL Queries Demonstrating Solution
- Nexstrain CSV — that includes the benefits of additional data enhancement from SPARQL INSERTS that integrate into the LOD Cloud Knowledge Graph via DBpedia Country Identifiers

SPARQL Results Page that includes Hyperlinks functioning as Super Keys used for Data Integration [Live Link]
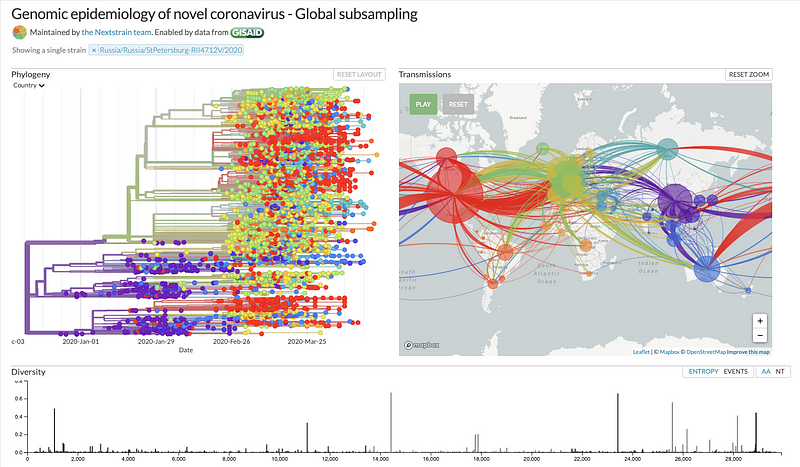
Nextstrain Page for a selected specimen submission
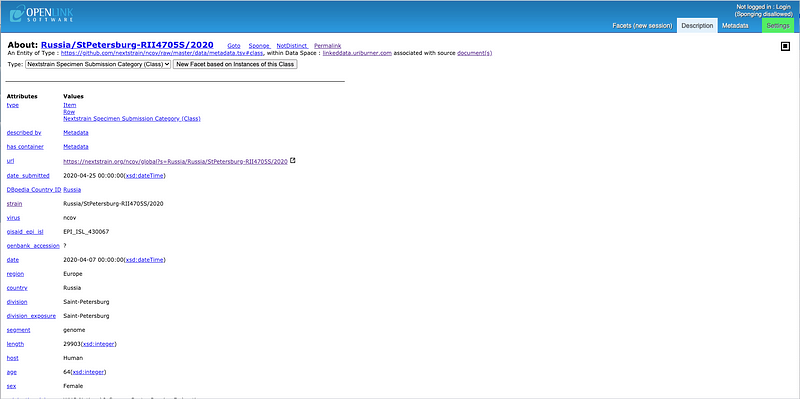
Specific Details about a SARS-CoV-2 Strain that also includes a bridge to the LOD Cloud KnowledgeGraph via DBpedia
- Johns Hopkins COVID-19 Daily Tracking — multiple CSV docs mapped and transformed to RDF that also includes benefits of additional data enhancements from SPARQL INSERTs that loosely-integrate with DBpedia and various Dashboards
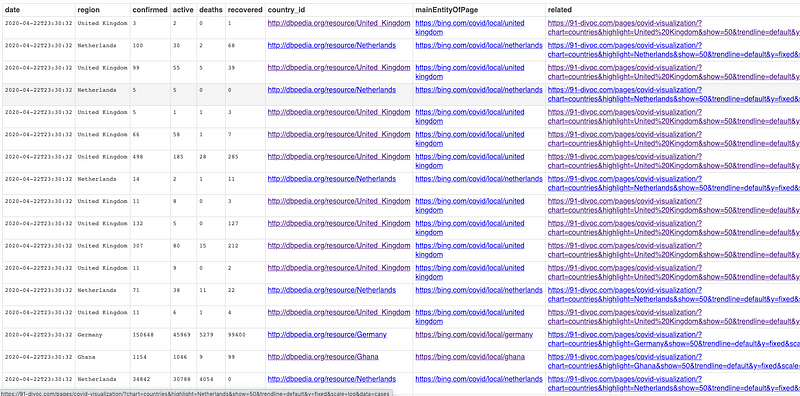
SPARQL Results Page that includes Hyperlinks functioning as Super Keys used for Data Integration [Live Link]
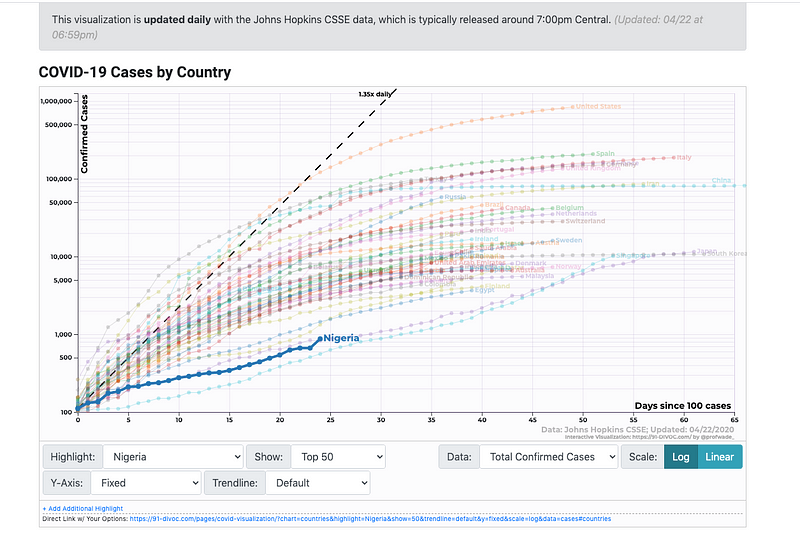
Divoc Dashboard scoped to Nigeria
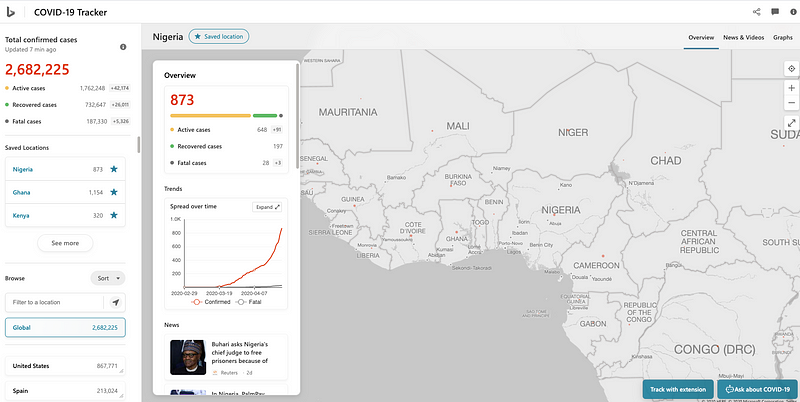
Bing Dashboard scoped to Nigeria
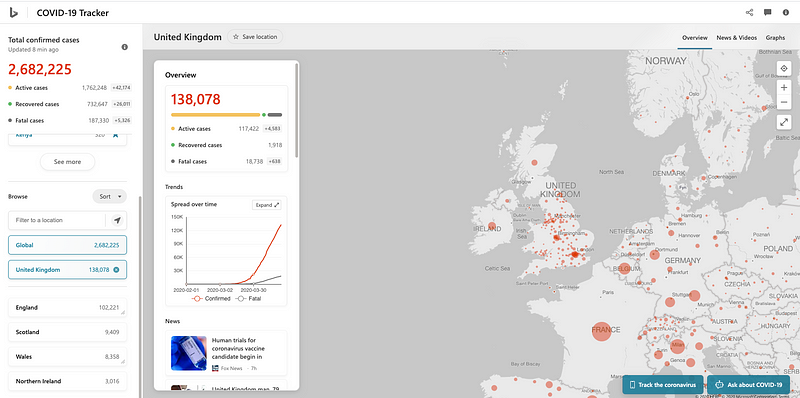
Bing Dashboard scoped to the United Kingdom
Conclusion
The COVID-19 pandemic is driven by a SARS-CoV-2 which like any virus seeks to exploit the vulnerabilities of its host — us the globally interconnected collective.
Change-sensitive access, integration, and management of disparate data sources is our greatest weapon against this and future outbreaks (which are guaranteed by the sheer mechanics of nature). That said, as we learned this time around, there is a massive price to pay if we don’t have the power of networks based on open standards at our disposal.
The Web and Internet networks made the initial shocks bearable, but its the emerging LOD Cloud Knowledge Graph (another network based on a Web of Data) that will eventually get us over the hump while providing critical foundation for newer networks of intelligence and beyond.
There is too much value to be created and exploited by future networks, so let’s not waste time and money on proprietary data access, integration, and management technology that operate strange business models based on ill-conceived notions that fuel the creation and perpetuation of Data Silos!
Related
- Master Google Spreadsheet for Open Data to Linked Open Data Transformation Templating
- Data Wrangling and Progressive LOD Cloud Knowledge Graph Updates — for COVID-19 and SARS-CoV-2 related Data Sources
- Nextstrain
- COVID-19 Tracking Project
- Johns Hopkins COVID-19 Daily Tracking Reports & Dashboards
- Our World In Data